By Xia Fan, science and Technology Daily Reporter
On April 8, the reporter learned from Zhejiang Wanli University in China that a team from the university’s School of Biology and Environment and teams from Hong Kong Baptist University and Ningbo Oriental University of technology had developed a high-precision sequencing technology called pNAD-seq, the highest resolution map of E. coli NAD-capped RNA was obtained, which provides a key methodological support for understanding the mechanism of this novel RNA modification in gene expression and environmental stress. The results were recently published in Nature Communications.
In eukaryotes, the 5′ end of the mRNA usually wears a“Cap”– 7-methylguanosine (M7G) , which not only protects the RNA from degradation, but also helps it translate into proteins. Scientists have long known that bacterial RNA has only bare triphosphate ends. In recent years, scientists have discovered that bacterial RNA can also wear another“Hat”-nicotinamide adenine dinucleotide (NAD +) .
NAD + is the core coenzyme of cellular energy metabolism, and also can be used as the initial nucleotide of RNA, which is“Sewn” to the RNA strand at the beginning of transcription to form NAD-RNA. However, due to technical bottlenecks, scientists have been difficult to accurately locate the origin of transcription, and it is difficult to effectively capture short NAD-RNA, resulting in undetected.
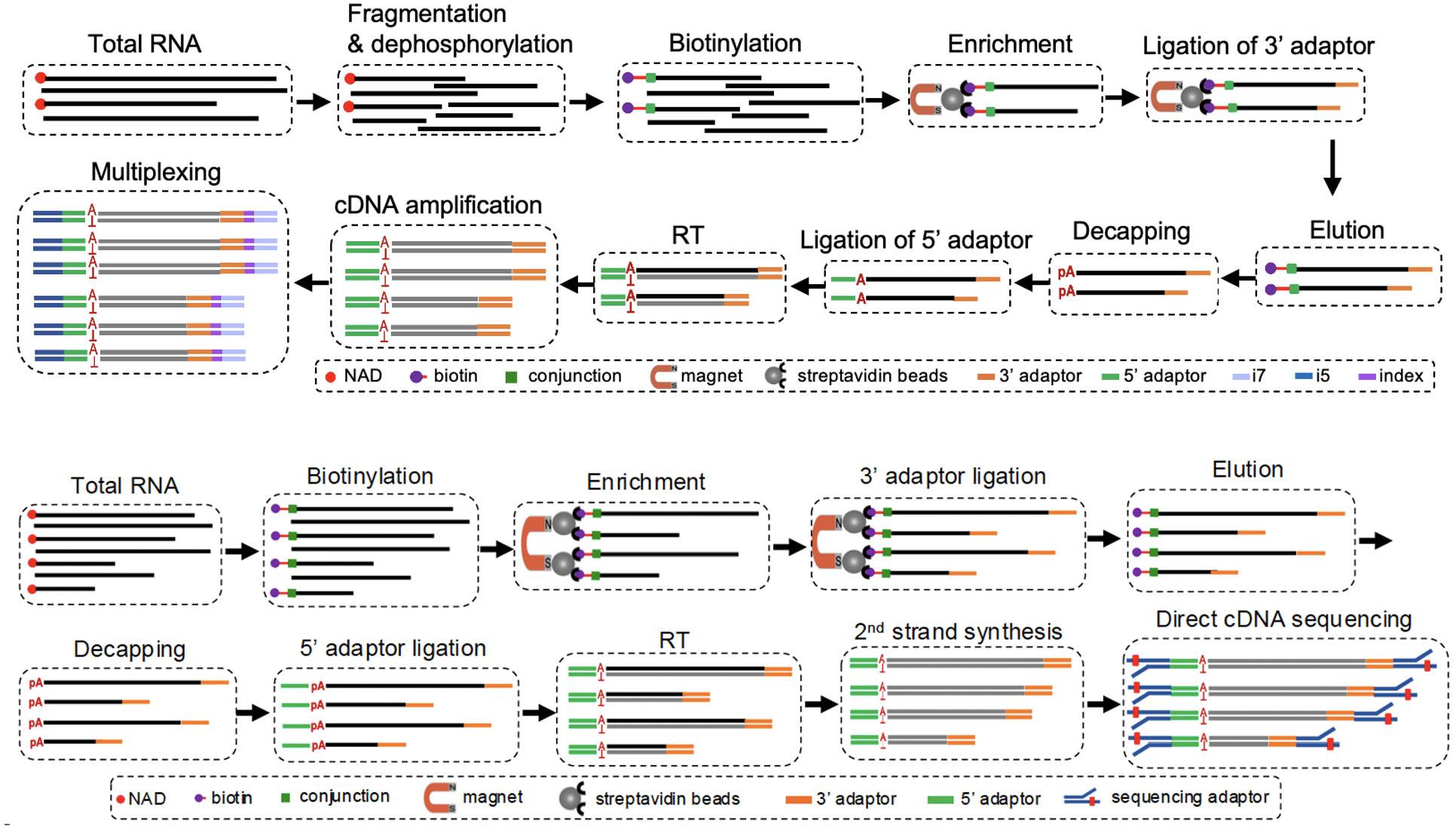
PNAD-SEQ (Top) and NAD linkSeq (bottom) flowcharts. (courtesy of respondent)
Dr. Zhang Hailei, from the School of Biology and environment of Zhejiang Wanli University, said that in order to see this“Hat” clearly, the team developed two complementary“Molecular trackers”: pnadseq, which uses ingenious chemical reactions to precisely capture and locate NAD-capped RNA; and NAD linkSeq, which uses the Nanopore sequencing platform, full-length sequences of these rnas are read.
With the help of the high-resolution tracker, the team has produced the most complete high-resolution map of the entire transcriptome of E. coli NAD-capped RNA to date. “Not only did it mark which genes were capped with NAD, but it also found a pattern of NAD capping: there was a mysterious conserved sequence, RDAY, near the transcription start site,” Zhang said, “It’s like finding a ‘code’ in the gene sequence and knowing it makes it possible to switch these genes on and off artificially in the future.”
It is understood that the research not only solves the long-standing technical pain points in the field, but also opens the door to explore the role of NAD capping in the pathogenicity, antibiotic resistance and synthetic biology of pathogens.